
Beck Agricultural Center is located NW of the Purdue University West Lafayette Campus on Highway 52. Directions are available here.
Registration covers both daily sessions (5/10-11), including the 5/10 poster session; no registration necessary for the 5/9 Public Kickoff. Light breakfast provided both days, as well as lunch and coffee breaks. Note: Attendance is in-person only – hybrid/online option not available.
Schedule
Day 1 (Tues, May 10)

(Click to enlarge)
Posters (Tues, May 10)
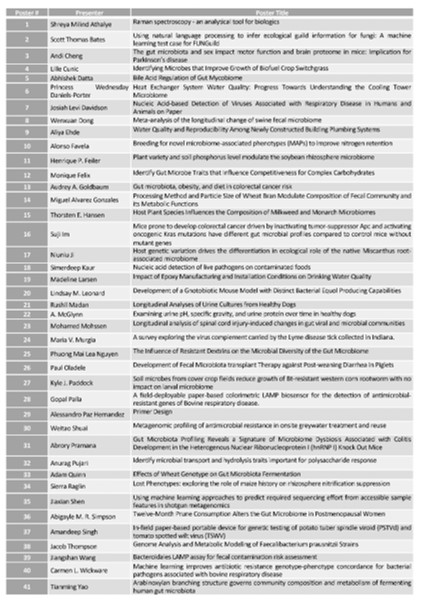
(Click to enlarge)
Day 2 (Wed, May 11)

(Click to enlarge)
Download/View Full Schedule/Program (including abstracts) HERE
Special Public Kickoff
James Hamblin
“Telling Stories about the Microbiome”
May 9, 2022 (7:00 PM)
Fowler Hall, Purdue University, West Lafayette, IN

Dr. James Hamblin is a preventive medicine physician, former staff writer at The Atlantic, and lecturer at Yale School of Public Health. He is the author of “If Our Bodies Could Talk” (Doubleday, 2016) and “Clean: The New Science of Skin” (Riverhead, 2020). He hosted the video series “If Our Bodies Could Talk”, for which he was a finalist in the Webby awards for Best Web Personality. A number of his topics focus on the microbiome, including personal microbiome testing, probiotics, and stopping showering.
His writing and videos have been featured in the New York Times, Washington Post, Los Angeles Times, Politico, NPR, The Guardian, Elle, Mother Jones, The Awl, and Marketplace, among others. Time named him among the 140 people to follow on Twitter, Greatist named him among the most influential people in health media, and BuzzFeed called him “the most delightful MD ever”.
Confirmed Speakers – Main Symposium
May 10-11, 2022
Beck Center, West Lafayette, IN
Greg Caporaso
Northern Arizona University

Dr. Greg Caporaso is an Associate Professor at Northern Arizona University, and Director of the Center for Applied Microbiome Science. Dr. Caporaso has published over 100 papers on the human microbiome, environmental microbiomes, and microbiome bioinformatics. He is the principal investigator on the popular QIIME microbiome bioinformatics platform (https://qiime2.org), a widely used tool in microbiome research that has been cited nearly 30,000 times since its publication in 2009, and the author of An Introduction to Applied Bioinformatics (http://readIAB.org).
Website: http://caporasolab.us
Amy Willis
University of Washington

Dr. Amy Willis is the Principal Investigator of the Statistical Diversity Lab and an Assistant Professor in the Department of Biostatistics at the University of Washington. Amy brings her expertise in statistical methods development, high dimensional data, statistical machine learning, phylogenetics and computational biology to develop tools for the analysis of microbiome and biodiversity data. She leads a team of statisticians creating methods and software to advance our understanding of microbial ecosystems, spanning a wide variety of application areas in human and environmental health. Her group works on what they believe to be the most critical methodological needs in microbial science, and the most serious shortcomings of existing analytical methods. She is also heavily involved in outreach, education, and collaboration as a core part of the SDL’s mission.
Website: http://statisticaldiversitylab.com/
Andy Benson
University of Nebraska-Lincoln

Located at the University of Nebraska Food for Health Center, Dr. Andy Benson’s research group studies the complex sets of host and dietary factors that collectively influence composition and function of the gut microbiome. In collaboration with statisticians, computational biologists, and animal geneticists, his research program has focused on understanding how individual genetics can influence the microbiome, and how dietary factors can modify the impact of host genetics, most likely through a direct impact of diet on the gut microbiome. Benson’s group is also spearheading the discovery component of the Nebraska Food for Health Center using complex trait analysis in crop plants to define components and molecules that can impact the gut microbiome of humans. Working closely with center members in plant genetics, statistics, and glycobiology, his team uses in vitro microbiomes in high-throughput screens of milled grains from large genetic resource populations of crop plants. This approach to complex phenotyping enables rapid and quantitative measurements of thousands of genetic variants to define pathways and molecules that are capable of influencing one or more members of the gut microbiome.
.Website: https://foodforhealth.unl.edu/andy-benson
Jennifer Pett-Ridge
Lawrence Livermore National Laboratory

Dr. Jennifer Pett-Ridge is a senior staff scientist and group leader at LLNL known for developing and applying isotopic tools that help us discover and quantify how changing climate shapes the roles of microorganisms and plants in environmental biogeochemical cycles. She has pioneered the use of NanoSIMS isotopic imaging in the fields of microbial biology and soil biogeochemistry, and in 2014 received a DOE Early Career award to work on responses of tropical soil microbes to climate change. As lead scientist of the LLNL DOE BER Genomic Science Biofuels Scientific Focus Area (SFA) from 2009–2018, and more recently the LLNL Soil Microbiome SFA (2018-present), she helps to coordinate multi-disciplinary teams that integrate biogeochemistry, stable isotope probing, NanoSIMS imaging, molecular microbial ecology, and computational modeling to understand biotic interactions and energy flow in microbial communities critical to soil nutrient cycling and sustainable biofuel production. She is the group lead for the Environmental Isotope Systems group in the Nuclear and Chemical Sciences division, the lead of LLNL’s National Getting to Neutral analysis of carbon sequestration capacity in the USA, and manages a portfolio of over $25 million in DOE, NSF, NASA, and other funding. She helps to mentor a group of staff scientists, postdocs, and graduate students working on terrestrial and marine carbon cycling, plant-soil interactions, and development of novel isotope tracing methods (El-FISH, Chip-SIP, STXM-SIMS) and collaborates frequently with scientists at academic institutions and other national labs. Pett-Ridge has published over 110 peer-review articles, including a patent ROI for the “ChipSIP” approach linking microbial identity and function using NanoSIMS analysis of microarrays.
.Website: people.llnl.gov/pettridge2